Training data balance is crucial to the performance of machine learning (ML) models, especially deep learning models. There would be a high risk of overfitting when training on unbalanced datasets.
Data insufficiency, however, is one of the biggest challenges of application of machine learning methods on melanoma detection. Deep Learning requires massive amounts of data. We will have massive amounts of lesions, but most of them will be benign, thus, our dataset will be highly unbalanced. In iToBoS project we will be collecting data from several hundreds of patients and some of them with a history of melanoma. However, there will be fewer melanoma cases than non-melanoma cases in the real world. Therefore, the number of melanoma images will be much less than other types of lesions. This will cause an unbalanced training dataset.
Using data augmentation techniques, we can add new data points to existing data to increase its amount artificially and balance the dataset. This may include making small changes to data or generating new data points with deep learning models. However, classical image processing techniques may not be good enough for augmentation of melanoma lesion images because they may cause complications in classification step or may not provide the desired variety that the model needs.
In iToBoS project we are trying to overcome melanoma case insufficiency by using generative adversarial networks (GANs) to create new synthetic data. GAN algorithms can learn patterns from input datasets and automatically create new examples which resemble training data. In our case we will train a GAN model with existing melanoma lesions captured by iToBoS scanner. When our model successes in learning skin images it would be capable of generating new realistic melanoma images and help us to have a balanced suitable dataset.
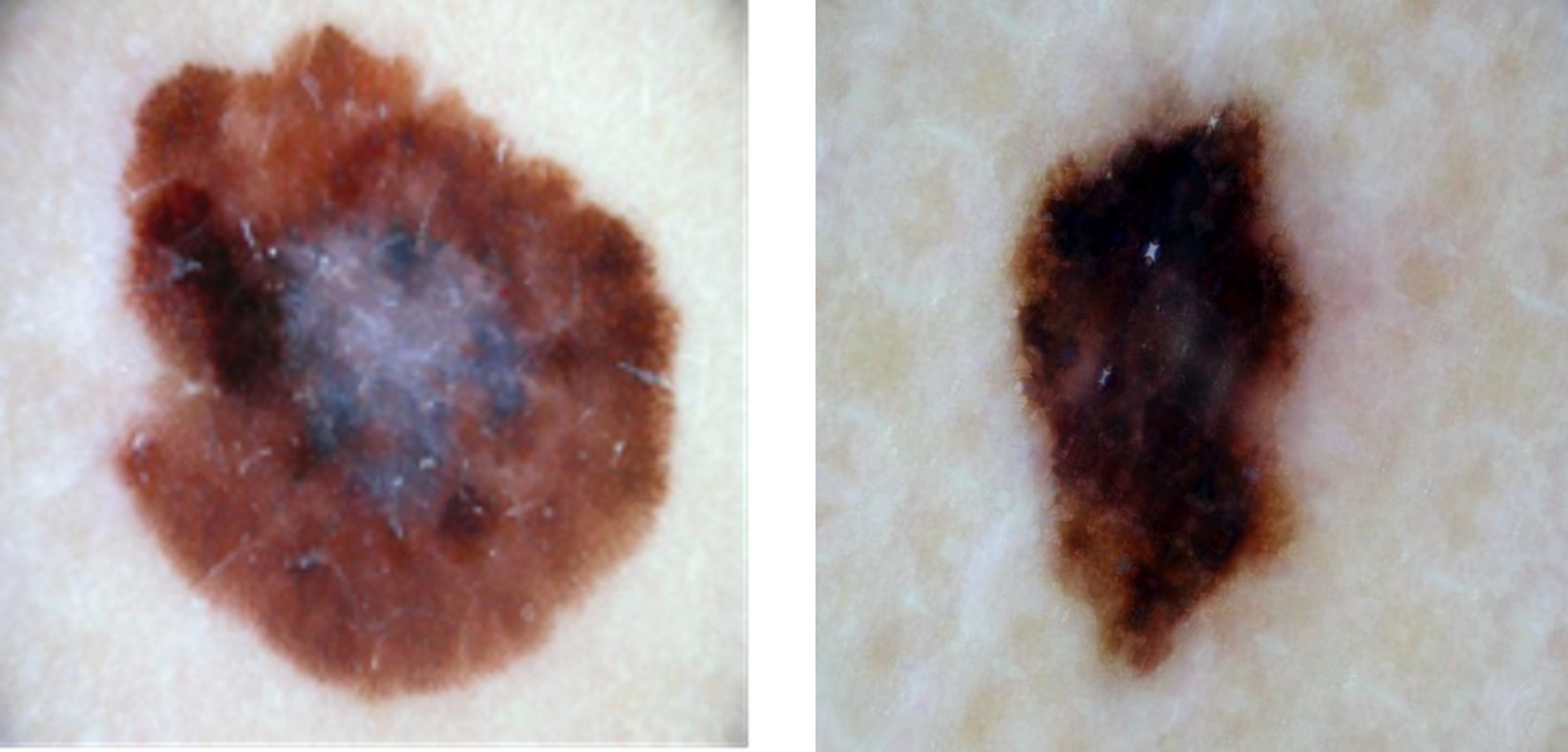
Sample real melanoma image from HAM10000 dataset (left); Sample fake melanoma image generated by our GAN, once trained using the HAM10000 dataset (right)
